Zebrafish larvae to test renal toxicity of an antibiotic
Renal toxicity is an important aspect to be assessed when a new drug is tested. To study this toxicity, in vitro cell-based assays are often used. Unfortunately, translating the results of such cell assays to vertebrates remains challenging. That is why scientists from the University of Basel have evaluated whether zebrafish larvae could serve as a vertebrate screening model to detect changes of renal organs induced by the antibiotic gentamicin.
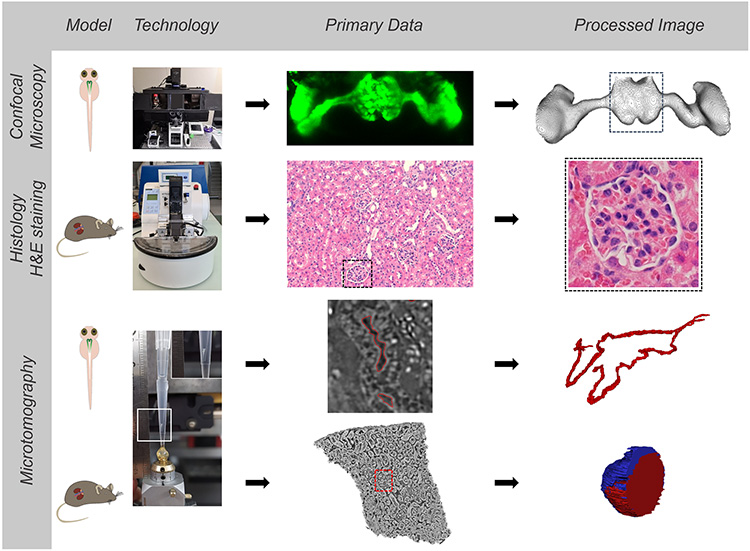
To visualize gentamicin-induced renal toxicity, they have used X-ray micro-computed tomography using synchrotron radiation (SRµCT) on the ANATOMIX beamline of SOLEIL. This method provides 3D representations of renal structures with micrometer resolution, without the need to label the organs studied.
Notably because the functional unit of their kidney closely resembles the human anatomy, zebrafish larvae are well suited to study renal morphological aberrations and are a predictive model for acute human kidney injury. Furthermore, this model organism is optically transparent during the early developmental stages and can be used as an alternative to animal experiments in higher vertebrates according to 3R principles of animal welfare*. Acute kidney injury in zebrafish can be induced by the use of nephrotoxins, for instance gentamicin, resulting in acute cellular damage of kidney tubules (i.e., the structures of zebrafish larvae shown in green in Figure 1). This includes hindered tubular development and their enlargement and necrosis.
Gentamicin is a drug that belongs to a class of antibiotics known as aminoglycosides. These are known to be toxic to the kidney and inner ear in humans. Gentamicin-induced renal morphological aberrations in kidneys are thus not specific to zebrafish larvae. The scientists from Switzerland assessed and monitored the impact of gentamicin on the kidney anatomy of zebrafish larvae and extrapolated the results to the kidney structure of mice. For their studies with visible-light microscopy, they used zebrafish lines genetically modified so that part of their renal structures (glomerulus) produced a green fluorescent protein (GFP), making it easier to visualize potential lesions of these structures when analyzed by confocal laser scanning microscopy.
But in the kidneys of gentamicin-treated zebrafish larvae, the fluorescent GFP signal disappears because of the nephrotoxic effects of the antibiotic. Therefore, the researchers also employed another analysis technique: SRμCT, implemented on SOLEIL's ANATOMIX beamline.
Gentamicin-treated and untreated zebrafish larvae and mice were imaged at ANATOMIX. Comparison of X-ray microtomography images revealed similar effects of the nephrotoxin in both species (Figure 1). In particular, the study revealed a similarity between morphological aberrations in zebrafish larvae and rodent kidney.
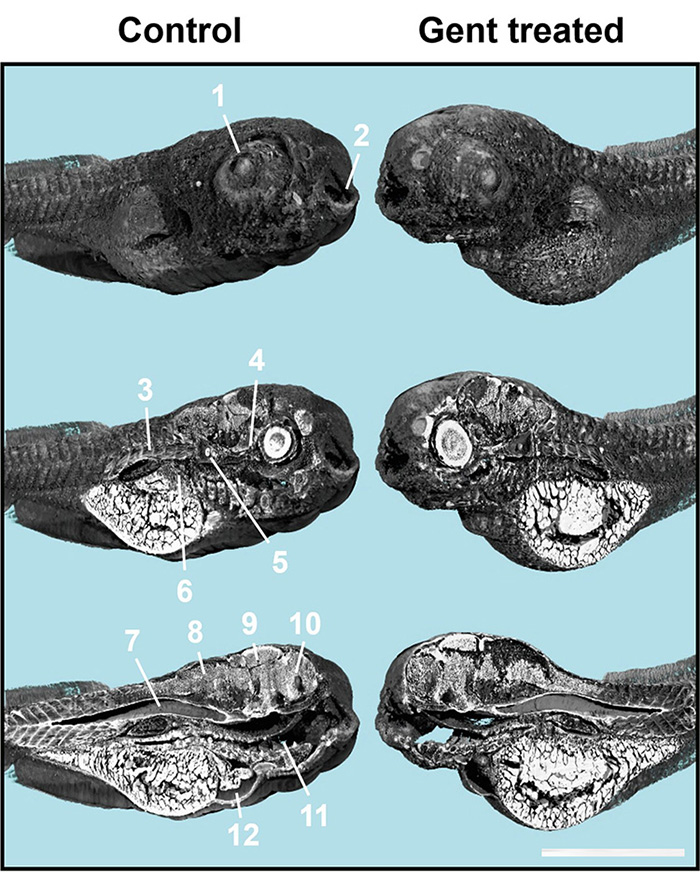
X-ray microtomography also allowed to obtain 3D images of the organs of non-genetically modified zebrafish larvae (that do not produce GFP, cf Figure 2) and of mice, down to the scale of individual cells.
1 = eye lens, 2 = mouth, 3 = muscle tissues, 4 = optical nerve, 5 = otoliths, 6 = liver, 7 = notochord, 8 = hindbrain, 9 = midbrain, 10 = frontbrain, 11 = intestine, 12 = heart.
The scientists conclude that zebrafish larvae may be suitable as an intermediate model to assist in the extrapolation from cell-culture-based test systems to mammals.
The approach can therefore be used to identify potential nephrotoxins in drug discovery or environmental sciences.
*3R principles: https://en.wikipedia.org/wiki/Three_Rs_(animal_research)