First biological application of the super-resolved infrared optical photothermal microscopy at the SMIS beamline
A scientific collaboration between Lund University (Sweden) and the SMIS beamline of Synchrotron SOLEIL elucidated for the first time the conformation of nanoscale amyloid aggregates. This study was conducted in mice, directly in neurons affected by Alzheimer's disease, using optical‐photothermal infrared microscopy, a new super-resolution technique combining a pulsed infrared light source and laser in the visible region.
Due to increased life expectancy and population aging, the prevalence of Alzheimer disease (AD) which represents 70% of senile dementia cases and affects 50 million people in the world, increases quickly. AD cannot be cured by currently available therapies, thus research in novel therapies and diagnostic means is necessary. Genetic, biological and pathological data support the causal role of the amyloid aggregation of the Aβ peptide in the development of the disease. The Aβ peptide is generated by cleavage of the APP protein. According to the ‘amyloid cascade’ theory, the gradual accumulation of monomeric Aβ peptides in neurons and the extracellular space lead to the oligomerisation and to the polymerization of Aβ monomers into amyloid fibrils and amyloid plaques and finally to cell death. Although Aβ monomers are considered to be innocuous, aggregation causes them to adopt the abnormal, neurotoxic β-amyloid conformation. Understanding the mechanism of β-amyloid aggregation is of primary importance to develop new treatments.
The aggregation mechanism can be studied in vitro but is expected to be different in vivo as reproducing neurons intracellular and extracellular microenvironments is difficult. Different methods can be used to visualize amyloid plaques in vivo (electon microscopy, confocal fluorescent microscopy with immunostaining, histological dyes) but they give little to no information on the peptide conformation, lack specificity and or require chemical processing that can alter peptide conformation.
Vibrational spectroscopy methods such as Fourier Transform Infrared (FTIR) is used to evaluate the secondary structure of peptides and proteins1,2, and the β-amyloid conformation exhibits a specific FTIR signature3,4. FTIR microspectroscopy (µFTIR) can detect β-amyloid in cells and in brain sections and give information on the conformation of monomers, oligomers, and higher order structures such as amyloid fibrils and plaques5. µFTIR spectroscopy at the SMIS beamline of Synchrotron SOLEIL was used to evidence the amyloid conformation of small, micron sized protein inclusions directly in neurons in histological sections from the brain of Huntington disease patients and mice models6,7. Even when using the synchrotron source, the spatial resolution achieved by µFTIR is limited by diffraction to a few microns (3-10 µm) and is wavelength dependent. In addition, spectral interpretation can be made impossible by the occurrence of Mie scattering, a specific type of scattering that occurs on circular objects with similar size than the wavelength, such as neurons in culture. This is due to the fundamental relationship between absorption and refraction (as expressed in the Kramers-Kronig law). Therefore, it is difficult to apply µFTIR to study the aggregation of Aβ directly in neurons in culture at subcellular resolution.
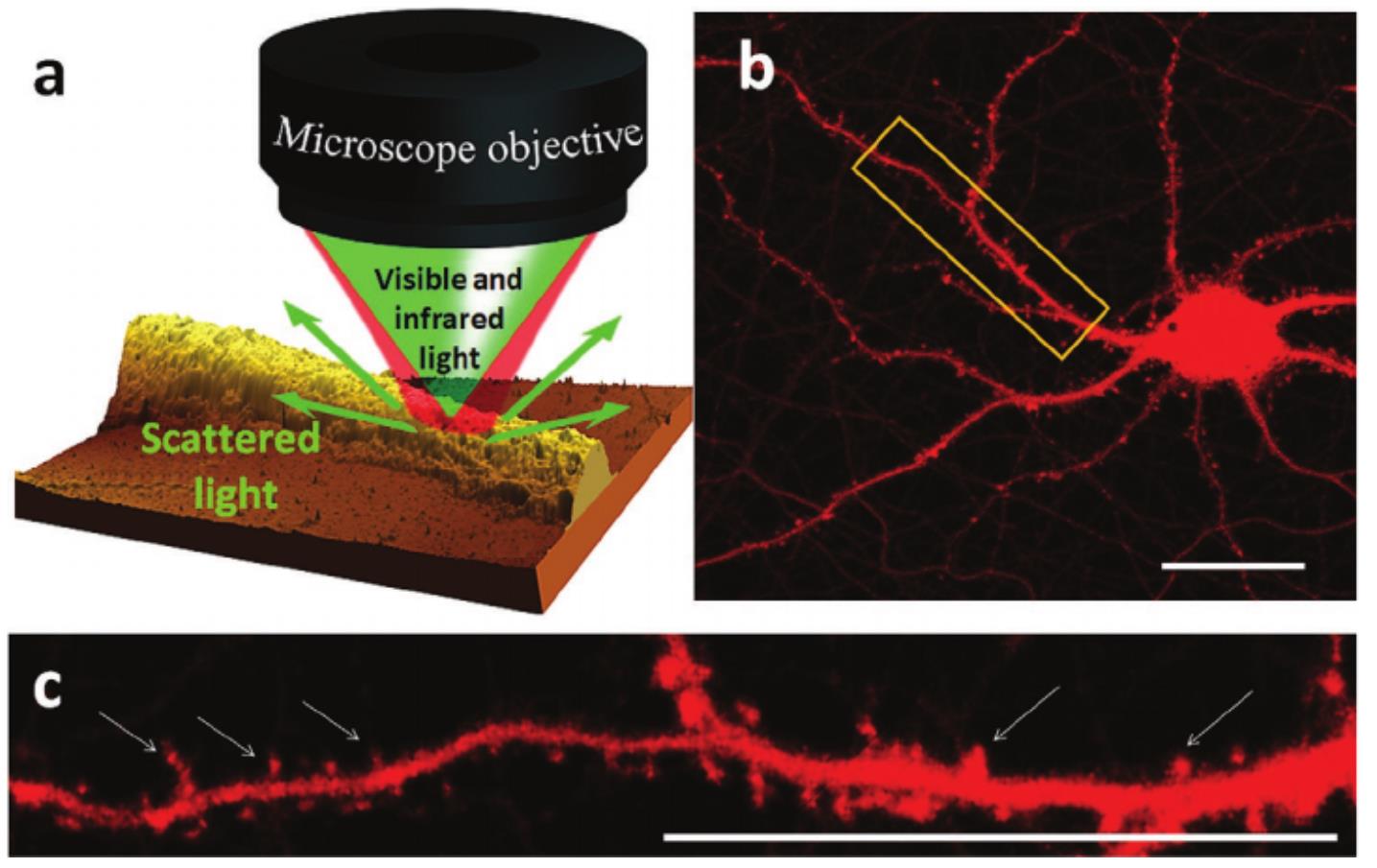
These limits can be overcome by using a new super-resolved technique called Optical Photo-Thermal IR spectroscopy (O-PTIR) which is available at the SMIS beamline since 2019. This technique uses the excitation of the sample by a brilliant pulsed IR source and the sample response is read by a laser in the visible domain (532 nm). It combines the chemical and conformation information from IR spectroscopy and the spatial resolution of visible microscopy and allows reaching spatial resolutions below the IR diffraction limit (400 nm in this study) without prior staining or chemical processing of the sample.
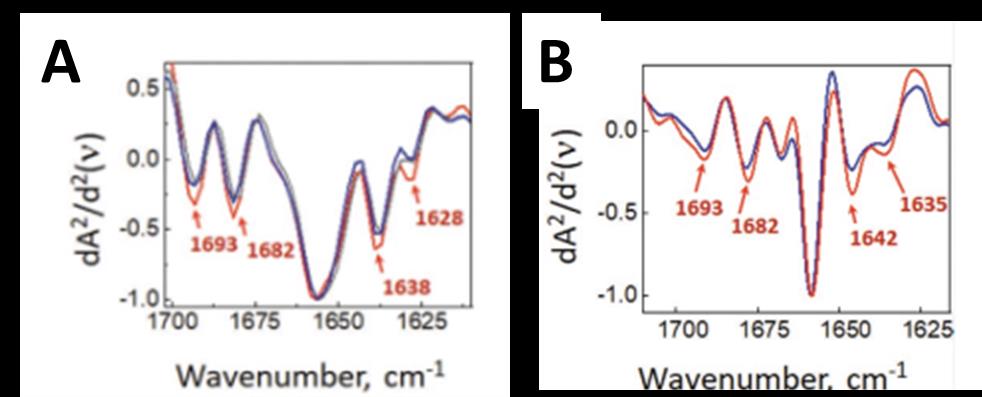
In this study8, a team of scientists from the SMIS beamline of Synchrotron SOLEIL and the team of Oxana Klementieva from the Lund University in Sweden studied the conformation of Aβ aggregates directly in neurons from genetically altered mice expressing the β1-42 fragment of human APP protein, model of Alzheimer's disease. The synchrotron µFTIR technique was used to measure the conformation of the Aβ aggregates at the scale of whole neurons and the O-PTIR technique for measuring the aggregates at subcellular resolution. Thanks to the high resolution of the O-PTIR technique, it was possible to detect β-amyloid aggregates a few hundred nanometers in size in neuron dendritic spines, which had never been achieved before. In addition, these nano-aggregates had a different spectral signature from that of the whole neuron revealing a polymorphism of the aggregates in vivo with slightly less ordered and probably non-fibrillar structure. The unordered β-sheet aggregates detected in diseased spine were structurally distinct from those found in neurites. This could correspond to the so-called amorphous amyloid aggregates which have been shown to be toxic to cells.
----------------------------
(1) Barth, A. Biochim. Biophys. Acta 2007, 1767 (9), 1073–1101
(2) Jackson, M.; Mantsch, H. H. Critical Reviews in Biochemistry and Molecular Biology 1995, 30 (2), 95–120
(3) Zandomeneghi, G. et al. Protein Sci. 2004 Dec; 13(12): 3314–3321
(4) Hiramatsu, H.; Kitagawa, T. Biochim. Biophys. Acta 2005, 1753 (1), 100–107
(5) Miller, L. et al. Biochim. Biophys. Acta 2013, 1828 (10), 2339–2346
(6) Hoffner, G. et al. Rev. Anal. Chem. 2014, 33 (4), 231–243
(7) André, W. et al. Anal. Chem. 2017, 89, 10, 5201–5209
(8) See "Related Publication" section beside