Cell signaling - Structure and Function of the C-terminal Domain of a Voltage-Gated Sodium Channel
Voltage-gated sodium channels are complex proteins present in the membranes of all higher organisms that are responsible for the transport of sodium ions into and out of cells. In humans, sodium channel mutations give rise to a number of neurological and cardiac diseases, and are associated with ageing and pain.
Britannic and American scientists have solved the complete 3D structure of a bacterial channel, which is a prototype for this kind of proteins. Thanks to these results, obtained notably via X-ray diffraction experiments carried out on PROXIMA1 beamline and published in Nature Communications, they propose a new mechanism of gating for the channel.
Voltage-gated sodium channels play essential roles in electrical signaling between cells because the first step of the action potential in excitable cells involves the opening of the sodium channel gate. In humans, sodium channel mutations give rise to a number of neurological and cardiac diseases, and are associated with ageing and pain; as a result these channels are the targets of many pharmaceutical drugs, including ones for treatment of epilepsy, chronic pain, and cardiovascular diseases. Because similar sodium channels are also present in insects, they are also the targets for agriculturally-important insecticides (including pyrethroids); crucially, however, insecticides must be able to distinguish between human and insect channels.
Sodium channels are also present in some bacteria, but those channels are slightly simpler (albeit with homologous sequences and features to eukaryotic channels); hence they are excellent for studying the structure and functioning of these proteins. Bacterial channel are tetramers consisting of transmembrane voltage-sensing and pore domains, and a cytoplasmic C-terminal domain (CTD). We have solved the first crystal structure of a sodium channel pore in the open state (McCusker et al, 2012), using data collected at the SOLEIL PROXIMA1 beamline.
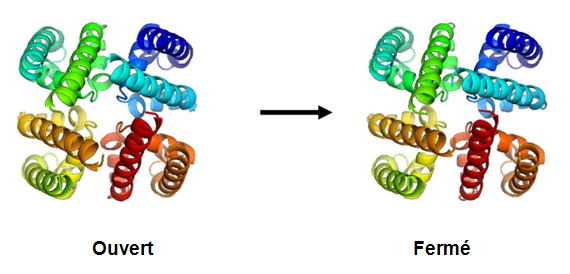
This structure enables us to understand how the channels open and close (by the rotation of a single bond in the middle of the S6 helix).
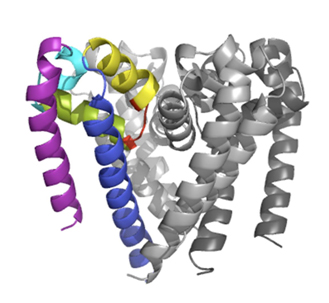
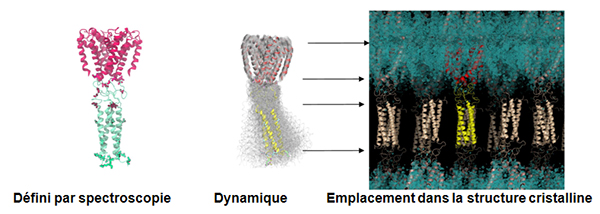
We have recently also solved a channel structure showing the location, structure and function of its C-terminal domain (Bagnéris et al, 2013) using the PROXIMA1 beamline. All previous crystal structures of bacterial sodium channels revealed the nature of their transmembrane domains but not their CTDs. We have used two spectroscopic methods, electron paramagnetic resonance spectroscopy combined with molecular dynamics (to determine its quaternary structure), and Synchrotron Radiation Circular Dichroism (SRCD) spectroscopic studies to determine its secondary structure (using data collected at the SOLEIL DISCO beamline – (Powl et al, 2010)) to show that the C-terminal domain of the NavMs channel from Magnetococcus marinus includes a flexible region linking the transmembrane domains to a four-helix coiled-coil bundle. Our new 2.9 Å resolution crystal structure of the NavMs pore depicts the CTD position, which is consistent with the spectroscopy-derived structures.
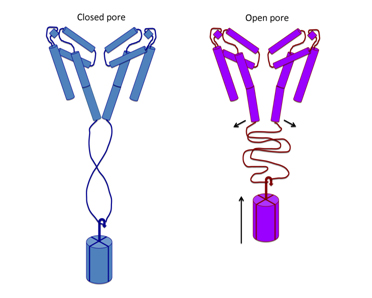
Electrophysiological studies of channel functioning demonstrate that the coiled-coil domain is important for channel stability and the flexible linking region couples channel inactivation with channel opening, and that negatively-charged residues in the linker region are essential for its function. A new mechanism for gating is proposed based on the structure, whereby splaying of the bottom of the pore is possible without requiring unravelling of the coiled-coil.
Références:
McCusker, E.C., Bagnéris, C., Naylor, C.E., Cole, A.R., D’Avanzo, N., Nichols, C.G., and Wallace, B.A. (2012) Structure of a bacterial voltage-gated sodium channel pore reveals mechanisms of opening and closing. Nature Communications 3:1102.
Powl, A.M., O’Reilly, A.O., Miles, A.J. and Wallace, B.A. (2010) Synchrotron radiation circular dichroism spectroscopy-defined structure of the C-terminal domain of NaChBac and its role in channel assembly. Proc. Natl. Acad. Sci. USA, 107:14064-14069.